
Differential expression of genes encoding sulfur metabolism-related periplasmic proteins of Acidithiobacillus ferrooxidans ATCC 23270
XIA Jin-lan(夏金兰), ZHANG Rui-yong(张瑞永), ZHANG Qian(张 倩), WU Shun(武 顺),
ZHANG Cheng-gui(张成桂), NIE Zhen-yuan(聂珍媛), QIU Guan-zhou(邱冠周)
Key Laboratory of Biometallurgy of Ministry of Education of China,
School of Minerals Processing and Bioengineering, Central South University, Changsha 410083, China
Received 11 September 2009; accepted 31 December 2009
Abstract: Reverse-transcription qualitative PCR (RT-qPCR) was used to analyze the changes in transcription levels of the sulfur metabolism-related periplasmic protein genes of Acidithiobacillus ferrooxidans ATCC 23270 grown on sulfur or ferrous. Seven periplasmic proteins with apparently higher abundance grown on elemental sulfur than on ferrous sulfate were identified by matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF-MS). Expression analysis of the corresponding genes by RT-qPCR shows that the constitutive expression of all those genes are more up-regulated grown on sulfur than those grown on ferrous (>10 folder). Study on the corresponding genes of the identified periplasmic proteins by RT-qPCR confirmed the results of two-dimensioned gel electrophoresis, indicating they may be related with sulfur metabolism in A. ferrooxidans.
Key words: Acidithiobacillus ferrooxidans; RT-qPCR; periplasmic proteins
1 Introduction
Elemental sulfur was produced during the dissolution of metal sulfides, which may be accumulated and form a sulfur layer onto the surface of minerals, leading to modification of the electrochemical properties of minerals surface and inhibition of the bioleaching process[1-2]. Actually, the elemental sulfur in bioleaching is basically chemical inert and it can be exclusively eliminated by sulfur-oxidizing bacteria[3-4].
The oxidation of elemental sulfur by Acidithiobacillus ferrooxidans (A. ferrooxidans for short) is a complex process which involves the attachment of cells to sulfur particles, the transport through the outer membrane and the oxidation. Most of the relevant reactions for sulfur oxidation take place at periplasmic space[5-6]. A thiosulfate sulfur-transferase (TST) of 21 kDa of A. ferrooxidans grown on elemental sulfur was found located in the periplasmic space and it broke the S―S bond present in thiosulfate, generating sulfur and sulfite[7]. Several other periplasmic proteins of A. ferrooxidans such as sulfur dioxygenase, sulfite oxidase and tetrathionate hydrolase were predicted to be involved in the oxidation of sulfur[8]. A total of 131 proteins in the periplasm of the microorganism grown on thiosulfate were identified by using high resolution linear ion trap-Fourier transform mass spectrometry (LTQ-FT/ MS)[9]. However, the mechanism of biological sulfur activation and oxidation in A. ferrooxidans is still unclear because of the lack of related researches all over the world.
In our previous study, the proteins present in the periplasmic fraction of A. ferrooxidans cultured on two different energy substrates, ferrous sulfate and elemental sulfur by two-dimensional gel electrophoresis were analyzed and characterized. To confirm the results of 2-DE and further study the genes that may be involved in sulfur oxidation, the expression of genes encoding periplasmic proteins with higher expression grown on sulfur were analysed by RT-qPCR. The results indicate that the expression of several genes encoding periplasmic proteins related to sulfur metabolism is induced when the microorganisms grow on element sulfur.
2 Experimental
2.1 Bacterial strain and growth condition
A. ferrooxidans ATCC 23270 was cultured in 9 K basal media supplemented with 5.0 g/L of elemental sulfur powder and 44.5 g/L of ferrous sulfate, respectively. The initial pH value of these media was adjusted to 2.0 with 1 mol/L H2SO4. The culture of A. ferrooxidans was grown in 500 mL flasks containing 300 mL medium and incubated at 30 °C with a drum pump to provide oxygen (10 L/min).
2.2 Preparation of periplasmic space proteins
A. ferrooxidans cells were resuspended in a solution (8 mL) containing 20% (w/v) sucrose, 33 mmol/L Tris-HCl (pH=8.0) and 0.5 mmol/L Na2EDTA. After shaking the incubation for 15 min, the shrunk cells were harvested by centrifugation at 8 000 r/min for 15 min. The supernatant was removed and the cell resuspended in 8 mL MiliQ water (4 °C). After shaking the incubation for another 10 min, the cell suspension was centrifuged at 10 000 r/min for 15 min, and then the periplasmic proteins were recovered in the supernatant.
2.3 Two-dimensional gel electrophoresis
Periplasmic proteins were separated by two- dimensional polyacrylamide gel electrophoresis (2-DE) performed with the IPGphor system and Ettan DALT II system (GE). After electrophoresis, proteins were visualized by Coomassie Brilliant Blue G-250 staining. Two-dimensional gels were scanned using a Uniscan scanner. The number of protein spots expressed on 2-DE was analysed by Imagemaster software version 7.2.
2.4 Microsequencing and MS analysis of proteins extracted from 2-D gels
Protein spots were excised and transferred into 1.5 mL microcentrifuge tubes and subjected to in-gel digestion. A total extract was concentrated in a vacuum centrifuge to 20 μL for further analysis. Peptide mass fingerprinting (PMF) were obtained by using an Autoflex MALDI-TOF/TOF mass spectrometer with delayed extraction and reflection[10-11].
2.5 RT-qPCR
A. ferrooxidans ATCC 23270 total cellular RNA was isolated using the TRIzol reagent (Invitrogen) according to the manufacturer’s protocol. RNA extracts were purified with an RNeasy mini kit (QIAGEN) according to the instructions, and the oncolumn DNase digestion was performed with RNase-free DNase (QIAGEN) to remove genomic DNA. Total cellular RNA was quantified at OD260 and OD280 with a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies). The purified RNA from each sample served as templates to generate cDNA.
RT-qPCR was performed with RNA extracts from samples grown on sulfur and ferrous. Total RNA ( l ?g ) was used to prepare cDNA by reverse transcription. They were denatured in the presence of 400 ng of the RT primer at 70 °C for 5 min and then cooled on ice. The RT reaction was carried out at 42 °C for 1 h in a final volume of 25 ?L according to the manufacturer’s instructions (Qiagen). After the RT reaction, the enzyme was inactivated by heating at 65 °C for 10 min, and a 10 ?L of the RT reaction was used for RT-qPCR.
The identified protein genes were detected by RT-qPCR. The primers used for RT-qPCR are described in Table 1. The RT-qPCR procedure was carried out according to Ref.[12]. The expression ratio was recorded as the fold difference in quantity of RT-qPCR product from samples grown at the treatment versus control concentration. Relative abundance of each gene versus a constitutively expressed gene (16S rDNA) was determined. The results were expressed as means ± standard errors of three independent experiments.
2.6 Sequence analysis
The functional categories for the different identified proteins were obtained from the genomic sequence of A. ferrooxidans (www.tigr.org). The presence of export signals for each protein found in the periplasmic fraction was predicted by using SignalP (www.cbs.dtu.dk/services/signalP), TatP (www.cbs.dtu.dk/ services/TatP-1.0), SecretomeP (www.cbs.dtu.dk/services/ SecretomeP) and LipoP (www.cbs.dtu.dk/services/ LipoP).
Table 1 Primers used for RT-qPCR

3 Results and discussion
3.1 Identification and predication of periplasmic proteins spots
As shown in Fig.1, there are some protein spots with relatively high abundance in the cells grown on sulfur as energy substrate but not on the ferrous sulfate as energy substrate. Seven protein spots with relatively high abundance on sulfur substrate are excised in gel. Identification and predication of these seven periplasmic proteins are made by MALDI-TOF/TOF MS followed by database searches against SwissProt and AFE-3217 based on the peptide mass fingerprints, and the results are shown in Table 2.
3.2 RNA isolation and expression studies by RT-qPCR
The quality of total cellular RNA: the ratios of OD260/OD280 quantified with a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies) was about 1.95-2.05. The integrality of total cellular RNA was checked by 1.5% agarose gel electrophoresis and ethidium bromide staining, as shown in Fig.2, which indicated that the purity and integrality of total cellular RNA was enough for this study.
To validate the results of 2-DE, RT-qPCR was used to analyze the changes in the transcription levels of the identified periplasmic protein genes of A. ferrooxidans ATCC 23270 grown on sulfur and ferrous. The results are given in Table 3. The mRNA expression of all the identified genes from the cells grown on sulfur was significantly higher than that from those grown on ferrous (>10 folder). One of the putative detoxification genes, AFE_3058 (encoding thio-specific antioxidant), was induced at the highest level, arrived 347-fold of those grown on ferrous, while the gene encoding AFE_2438 (cation ABC transporter) was induced at the lowest level, it still expressed 15.4-fold higher, when grown on sulfur.

Fig.1 Periplasmic proteins from cells grown on sulfur (a) or on ferrous iron (b) separated by 2-D PAGE
In the current study, the identified periplasmic proteins were further grouped into different categories according to their potential biological functions in the cell, as shown in Table 2.
Table 2 Identification and predication of periplasmic proteins spots of A. ferrooxidans grown on sulfur energy substrate by MALDI-TOF/TOF MS followed by database searches against SwissProt and AFE-3217 based on peptide mass fingerprint
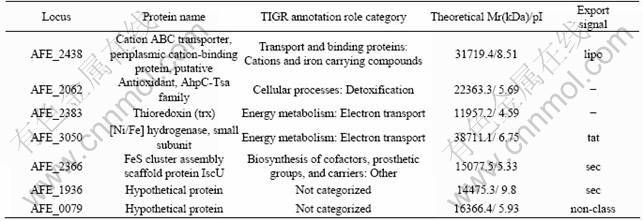
Table 3 Selected Acidithiobacillus ferrooxidans ATCC 23270 genes up-regulated grown on sulfur
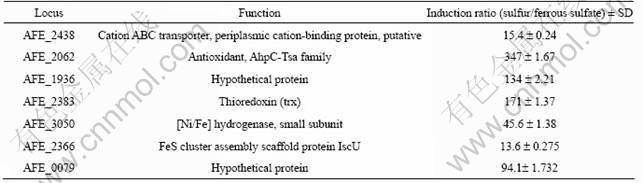

Fig.2 Electrophoresis of total RNA
A number of transport and binding genes appear to play roles in sulfur metabolism of A. ferrooxidans. In the present study, the expression level of transport and binding protein encoding gene AFE_2438 (cation ABC transporter) was up-regulated when grown on sulfur (15.4 fold), which was in accordance with the 2-DE. AFE_2438 is a putative periplasmic cation binding protein of the TroA family. Downstream of the genetic context of AFE_2438, there are genes coding for proteins with similarities to the high affinity of zinc ABC-type transporter: the ATP-binding protein (AFE_2437) and a duplicated permease (AFE_2436 and AFE_2435) that could form part of one transcriptional unit.
As an important model organism for bioleaching and bioremediation studies which could oxidize sulfur and ferrous, the oxidation process which takes place at the periplasm space is readily impacted by energy substrate. Thus, it is not surprising that a number of cellular processes and energy metabolism genes show substrates dependent expression in our study. All identified genes assigned to these two functional categories exhibit significantly up-regulated expression when grown on sulfur. Genes encoding putative antioxidant (AFE_2062), thioredoxin (AFE_2383) and [Ni/Fe] hydrogenase (AFE_3050) play a 45.6-347 fold range grown on sulfur than grown on ferrous. AFE_2062 was a putative antioxidant belonging to AhpC-TSA ( thiol specific antioxidant ) family which is functionally predicted to take part in the detoxification of cells, such as protection of cellular components against oxidative damage from a system capable of generating reactive sulfur species[13], while AFE_2383 was found to be a thioredoxins (trx) homologue which acted as antioxidants by facilitating the reduction of other proteins by cysteine thiol-disulfide exchange[14]. On the other hand, AFE_3050 appeared to be a bifunctional enzyme endowed with both proton- and sulfur-reducing activities[15]. The up-regulated genes encoding cellular processes and energy metabolism proteins grown on sulfur suggested that these genes were related with sulfur metabolism in this acidophile.
Iron-sulfur clusters are one of the most ancient and ubiquitous redox centers in almost all living organisms[16-17]. Biochemical studies indicated that iron-sulfur cluster assembly scaffold protein (IscU) is essential for the general biogenesis of iron-sulfur proteins in bacteria[18]. In the study, AFE_3058 (IscU) was detected high expression when grown on sulfur by 2-DE, which may be related to a large number of synthesis of iron-sulfur proteins. In addition, the expression of this differentially expressed gene was confirmed by RT-qPCR, suggesting that the transcription of IscU (AFE_3058) was increased approximately 13.6-fold grown on sulfur compared to those grown on ferrous.
Two differentially expressed unclassified functional genes in A. ferrooxidans are detected. Although the functions of these genes are not clearly determined, it is assumed that they may play potential roles in sulfur metabolism in A. ferrooxidans. In further study, RNA interference (RNAi) or some other techniques shall be used to verify the functions of the differentially expressed genes.
4 Conclusions
1) The proteins present in the periplasmic fraction of A. ferrooxidans cultured on two different energy substrates, ferrous sulfate and elemental sulfur were separated by two-dimensional gel electrophoresis. A total of seven periplasmic proteins with apparent higher abundance grown on element sulfur than on ferrous sulfate were identified by MALDI-TOF-MS.
2) The corresponding genes of the identified periplasmic proteins were further researched by RT-qPCR. All these genes were more significantly up-regulated grown on sulfur than those grown on ferrous (>10 folder).
3) The researches of the corresponding genes of the identified periplasmic proteins by RT-qPCR confirm the results of 2-DE, indicating they may be related with sulfur metabolism in A. ferrooxidans. The functions of the differentially expressed genes need to be further researched.
References
[1] SCHIPPERS A, SAND W. Bacterial leaching of metal sulfides proceeds by two indirect mechanisms via thiosulfate or via polysulfides and sulfur [J]. Applied and Environmental Microbiology, 1999, 65(1): 319-321.
[2] ROHWERDER T, GEHRKE T, KINZLER K, SAND W. Bioleaching review (part A): Progress in bioleaching: fundamentals and mechanisms of bacterial metal sulfide oxidation [J]. Applied Microbiology and Biotechnology, 2003, 63(3): 239-248.
[3] KNICKERBOCKER C, NORDSTROM D K, SOUTHAM G. The role of "blebbing" in overcoming the hydrophobic barrier during biooxidation of elemental sulfur by Thiobacillus thiooxidans [J]. Chemical Geology, 2000, 169(3/4): 425-433.
[4] ROHWERDER T, SAND W. The sulfane sulfur of persulfides is the actual substrate of the sulfur-oxidizing enzymes from Acidithiobacillus and Acidiphilium spp [J]. Microbiology, 2003, 149(1): 1699-1710.
[5] RAWLINGS D E. Characteristics and adaptability of iron- and sulfur-oxidizing microorganisms used for the recovery of metals from minerals and their concentrates [J]. Microbial Cell Factories, 2005, 4(1): 13-27.
[6] VALENZUELA L, CHI A, BEARD S, ORELL A, GUILIANI N, SHABANOWITZ J, HUNT D F, JEREZ C A. Genomics, metagenomics and proteomics in biomining microorganisms [J]. Biotechnology Advances, 2006, 24(2): 197-211.
[7] TABITA R, SILVER M, LUNDGREN D G. The rhodanese enzyme of ferrobacillus ferrooxidans (Thiobacillus ferrooxidans) [J]. Biochemistry and Cell Biology-Biochimie Et Biologie Cellulaire, 1969, 47(12): 1141-1145.
[8] RAMIREZ P, TOLEDO H, GUILIANI N, JEREZ C A. An exported rhodanese-like protein is induced during growth of Acidithiobacillus ferrooxidans in metal sulfides and different sulfur compounds [J]. Applied and Environmental Microbiology, 2002, 68(4): 1837-1845.
[9] CHI A, VALENZUELA L, BEARD S , MACKEY A J, SHABANOWITZ J, HUNT D F, JEREZ C A. Periplasmic proteins of the extremophile Acidithiobacillus ferrooxidans [J]. Molecular and Cellular Proteomics, 2007, 6: 2239-2251.
[10] HANNA S L, SHERMAN N E, KINTER M T, GOLDBERG J B. Comparison of proteins expressed by pseudomonas aeruginosa strains representing initial and chronic isolates from a cystic fibrosis patient: an analysis by 2-D gel electrophoresis and capillary column [J]. Microbiology, 2000, 146(1): 2495-2508.
[11] SRISOMSAP C, SAWANGAREETRAKUL P, SUBHASITANONT P, PANICHAKUL T, KEERATICHAMROEN S, LIRDPRAPAMONGKOL K, CHOKCHAICHAMNANKIT D, SIRISINHA S, SVASTI J. Proteomic analysis of cholangiocarcinoma cell line[J]. Proteomics, 2000, 4(4): 135-1144.
[12] CHAO Jing,WANG Wei,XIAO Shui-ming, LIU Xue-duan. Response of Acidithiobacillus ferrooxidans ATCC 23270 gene expression to acid stress [J]. World Journal of Microbiology and Biotechnology, 2008, 24(10): 2103-2109.
[13] KIM K, KIM I H, LEE K Y, RHEE S G, STADTMAN E R. The isolation and purification of a specific "protector" protein which inhibits enzyme inactivation by a thiol/Fe(III)/O2 mixed-function oxidation system [J]. Journal of Biological Chemistry, 1988, 263(10): 4704-4711.
[14] AGO T, SADOSHIMA J. Thioredoxin and ventricular remodeling [J]. Journal of Molecular and Cellular Cardiology, 2007, 41 (5): 762-773.
[15] BRUGNA M, NITSCHKE W, TOCI R, BRUSCHI M, GIUDICI-ORTICONI M T. First evidence for the presence of a hydrogenase in the sulfur-reducing bacterium Desulfuromonas acetoxidans [J]. Journal of Bacteriology, 1999, 181(17): 5505-5508.
[16] KILEY P J, BEINERT H. The role of Fe-S proteins in sensing and regulation in bacteria [J]. Current Opinion in Microbiology, 2003, 6(2): 181-185.
[17] REES D C, HOWARD J B. The interface between the biological and inorganic worlds: iron-sulfur metalloclusters [J]. Science, 2003, 300(5621): 929-931.
[18] YANG J J, JACOB P B, DINGH G. Interplay of IscA and IscU in biogenesis of iron-sulfur clusters [J]. Journal of Biological Chemistry, 2006, 281(9): 27956-27963.
(Edited by FANG Jing-hua)
Foundation item: Projects(50674101, 50974140) supported by the National Natural Science Foundation of China; Project(2010CB630902) supported by the National Basic Research Program of China
Corresponding author: XIA Jin-lan; Tel: +86-731-88836944; E-mail: jlxia@mail.csu.edu.cn
DOI: 10.1016/S1003-6326(10)60656-5