J. Cent. South Univ. Technol. (2009) 16: 0416-0421
DOI: 10.1007/s11771-009-0070-y
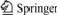
Isolation, identification and characterization of
cadmium-resistant Pseudomonas aeruginosa strain E1
ZENG Xiao-xi(曾晓希)1, 2, TANG Jian-xin(汤建新)2, LIU Xue-duan(刘学端)1, JIANG Pei(蒋 佩)2
(1. School of Resources Processing and Bioengineering, Central South University, Changsha 410083, China;
2. College of Packaging and Printing, Hunan University of Technology, Zhuzhou 410083, China)
Abstract: Strain E1 with resistance to 18 mmol/L cadmium (Cd), isolated from Cd-contaminated soil was identified by morphological observation, biochemical and physiological characterization and 16S rDNA sequence analysis. The resistance to heavy metals Cd, Cu, Co, Mn, Pb, Zn and 12 antibiotics was examined. The ability of removing Cd from solution was studied. The characterizations show that strain E1 is affiliated to Pseudomonas aeruginosa (P. aeruginosa). Strain E1 has high resistance to heavy metals and the order is found to be Cd>Mn>Zn>Cu>Pb>Co in solid media. Strain E1 also exhibits the resistance to 12 antibiotics. Both living and non-living cells of strain E1 can remove Cd from solution, and living cell has better biosorption than non-living cell.
Key words: identification; P. aeruginosa; cadmium-resistance; screening
1 Introduction
Cadmium (Cd) is widely applied in industrial processes including electroplating, plastics, batteries, alloy manufacturing and paints, and released from mining and metallurgical processes as a byproduct, resulting in the increasing environmental pollution [1]. As a heavy metal, cadmium has fatal effects on organisms and is extremely toxic at relatively low dosage [2]. Cd can transfer and accumulate in plants, animals and humans by food chains, leading to serious damage to human’s health [3]. Cd affects cell proliferation, differentiation, apoptosis and increases oncogene activation to carcinogenesis [4-5]. Cd exposure is associated with organism aging, peripheral arterial disease, vertebral osteoporosis and fractures, toxicity to neuron [6-9]. Cd selectively accumulates in lung, renal, liver, pancreas, bone and damages them [10-11]. NISHIJO et al [12] performed a 15-year follow-up study with the inhabitants living in the Cd-polluted Kakehashi River Basin in Japan. The results showed that renal tubular dysfunction induced by Cd affected the causes of death, and mortality for heart failure, cerebral infarction, and nephritis and nephrosis was increased among inhabitants living in a Cd-polluted area.
The environmental pollution caused by heavy metals has received increasing attention from the worldwide. Removal of heavy metals from the contaminative soil and water is crucial for environmental protection. The treatments of heavy metals removal from wastewater mainly include chemical precipitation, ion exchange, membrane technology and biosorption [13-14]. Compared with other methods, biosorption with the advantages such as low cost, without secondary pollution and facility has been received much attention [15].
Microorganisms in soil are sensitive to the high concentrations of heavy metals (Zn, Mn, Co, Cu) and the unnecessary heavy metals (Cd, Hg, Ag) at low concentration [16]. Microorganisms adapt to the heavy metal-polluted soil by changing intrinsic biochemical and structural properties, physiological, and genes. Therefore, the survival of microbes in polluted soil are prone to show higher resistance to heavy metals as compared to populations in non-contaminated sites [17]. Some heavy metal-resistant microorganisms isolated from polluted soils have shown the capacity of biosorption [18-20]. So, isolation of heavy metal-resistant bacteria is important to improve the application of microorganism to environment protection.
In this work, a Cd-resistant strain was isolated from Cd-contaminated soil, and identified as P. aeruginosa based on morphological observation, biochemical and physiological characterization, and 16S rDNA sequence analysis. The resistance to other heavy metals (Cu, Co, Mn, Pb and Zn) and the antibiotic was examined. Furthermore, the cadmium removal capacity was studied.
2 Experimental
2.1 Isolation of cadmium-resistant bacteria
The soil samples were collected from the sites which had piled up with cadmium mass for many years in Zhuzhou Smelter, Hunan Province, China. 0.2 mol/L stock solution of cadmium was prepared by dissolving CdCl2 in deionized water and sterilized by using 0.22 μm pore-size sterile filters. Beef extract-peptone medium was as follows: beef extract 3.0 g/L, peptone 10.0 g/L and NaCl 10.0 g/L.
10 g soil sample was added to 90 mL sterilized water, and mixed on the magnetic blender for 30 min to separate bacteria from the soil completely. After being deposited for 20 min, 1 mL suspension was added to liquid medium with 2 mmol/L Cd, incubated at 30 ℃ and 180 r/min in a rotary shaker for 3-4 d. The culture was plated onto the solid medium with 2 mmol/L Cd, and the colonies were streaked on solid media ranging from 3 to 19 mmol/L Cd to examine Cd-tolerance.
The microorganism that could tolerate the highest Cd concentration was selected and used in subsequent experiments. Single colonies were picked and streaked on the solid medium. Streak the plate for more than three times to get pure bacterium repeatedly.
2.2 Characterization of bacteria
The morphological characterizations of colonies growth were observed on agar plate. Grams staining, flagella staining, capsule staining, and spore staining of bacteria in the exponential growth phase were performed. And the bacteria were observed under the microscope. According to Ref.[21], some physiological and biochemical experiments were carried out.
For further identification, the genomic DNA was extracted using Genomic DNA Purification Kit according to the manufacturer’s instructions, and the 16S rDNA gene was amplified by PCR using the universal 16S rDNA primers 27f (5’-GAGAGTTTGATCCTGGCTCAG) and 1521r (5’-AAGGAGGTGATCCAGCC) [22]. The reaction conditions were as follows: 95 ℃ for 3 min; 32 cycles: 94 ℃ for 1 min, 56 ℃ for 1 min and 72 ℃ for 2 min; 72 ℃ for 10 min and 4 ℃ pause. PCR products corresponding to the expected size of amplified 16S rDNA (1.5 kb) were purified with a Gel Extraction Kit and sent to Shanghai Sangon Biotechnologies Co. Ltd. for sequencing.
16S rDNA sequence was submitted to the database of GenBank and compared with similar sequences by BLAST analysis. Phylogenetic tree of 16S rDNA was constructed using the soft ware Clustal W.
2.3 Resistance to other heavy metals
The minimal inhibitory concentration (MIC) means the lowest concentration of metal that completely prevents bacterium growth. 0.2 mmol/L stock solution of the other heavy metals including CuCl2, CoCl2, ZnSO4, MnSO4 and Pb(Ac)2 was prepared by the same manner of Cd solution. The agar plate method was used. Beef extract-peptone agar plates were supplemented with different concentrations of the five metals. The strain was streaked on the above plates. The plates were inoculated at 30 ℃ for 3-5 d to determine the MIC of each heavy metal. The control experiment was carried out using E .coli stored in the laboratory.
2.4 Antibiotic susceptibility
The antibiotic disks were purchased from Hangzhou Microbe Reagent Limited Company, China. The antibiotic disks contained 12 antibiotics: penicillin (10 IU), ampicillin (10 μg), amikacin (30 μg), streptomycin (10 μg), neomycin (30 μg), tetracycline (30 μg), erythromycin (15 μg), kitasamycin 15 μg), nalidixic acid (30 μg), polymyxin (300 IU), novobiocin (30 μg), and chloramphenicol (30 μg). The strain was tested for its sensitivity to the 12 antibiotics. 0.1 mL culture was plated onto beef extract-peptone agar plates. The antibiotic disks were placed on the plates and incubated at 30 ℃ for 3-5 d.
2.5 Cadmium removal
The cadmium removal capacity of the strain was tested preliminarily with shake flasks. Nine 250-mL shake flasks with 100 mL beef extract-peptone liquid medium were marked as No.1-9, and grouped as follows: (1) group A: No.1-3; (2) group B: No.4-6; (3) group C: No.7-9. The 3 flasks of group A with no-inoculation, the other 6 flasks with 1% inoculation, and the nine flasks were put into the rotary shaker at 30 ℃ and 180 r/min. Culturing for 24 h, the three flasks of group C were sterilized at 121 ℃ for 10-15 min and almost all the bacteria were completed under microscope. Then CdCl2?6H2O was added into the nine flasks to obtain final Cd concentration of 1 mmol/L. After the nine flasks were inoculated in rotary shaker for 24 h, the cultures from each flask were centrifuged at 8 000 r/min for 10 min and the cadmium concentrations of the supernatants were determined using an atomic absorption spectrophotometer. The average value was obtained and put into the following formula to gain the cadmium removal rate (ηCd).
ηCd=[(a-b)/a]×100% (1)
where a is the mass of cadmium in the solution without bacteria, and b is the mass of cadmium in the solution with bacteria.
3 Results and discussion
3.1 Bacterium isolation
There were a few small colonies on the agar plate with 18 mmol/L Cd after incubating for 2-3 d. When the Cd concentration increased to 18.5 mmol/L and incubated for 7 d, there were no colonies on the plate. The colony was streaked on the agar plate to get the purified bacterium and denoted as strain E1.
The colonies are lichen, irregular, glossy, wet and viscous. They can produce blue or blue-green pigment in beef extract-peptone medium. The bacterium is Gram-negative and has polar flagellum, without capsule and spore. It can move fast under optical microscope observation. The SEM observation shows that it is rod-shaped, blunt-round in both ends and the size is (0.5-1.0) μm×(2.0-3.0) μm (Fig.1). When it is cultured in 3 mmol/L Cd medium, its size becomes smaller to (0.3-0.6) μm×(1.1-1.5) μm (Fig.2). Cd is toxic to bacteria and prevents them from growing, so the size is smaller. Moreover, the smaller size can reduce the surface area and the mass of cadmium in the cell, resulting in lower toxicity to bacteria. This may be a self-protection of the strain to the poor environment.
Blue pigment is a characterization of P. aeruginosa. In media without Cd, strain E1 can produce blue pigment. When the Cd concentration is up to 1 mmol/L, the
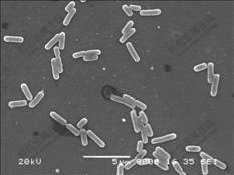
Fig.1 SEM image of strain E1 cultured in medium without Cd

Fig.2 SEM image of strain E1 cultured in medium with 3 mmol/L Cd
pigment disappears. It can be seen that Cd can inhibit the pigment formation.
Physiological and biochemical characteristics of strain E1 are listed in Table 1. According to Ref.[21], comparing strain E1 with other bacteria, the strain is similar to P. aeruginosa.
Table 1 Characteristics of strain E1
16S rDNA sequence of strain E1 is submitted to the database of GenBank and the submission number is EU375847. The BLAST analysis shows that the partial 16S rDNA of E1 is more than 98% identical with that of P. aeruginosa. In order to determine the relationship between strain E1 and the other P. aeruginosa, the phylogenetic tree based on 16S rDNA was constructed (Fig.3).
Based on the results of morphologic characteristics, physiological and biochemical characteristics and 16S rDNA sequence analysis, strain E1 is identified as P. aeruginosa.
3.2 MIC of other heavy metals
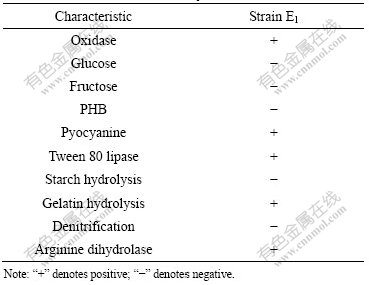
Strain E1 is able to grow at high concentrations of Cd, Cu, Co, Mn, Pb and Zn media. Cu, Co, Mn and Zn are essential to bacterium as trace nutrients. The others Cd and Pb have fatal roles. Heavy metals are toxic at micromolar or milimolar concentration. Table 2 lists the MIC of Cd, Cu, Co, Mn, Pb and Zn for strain E1 and E.coli. It can be seen that strain E1 can resist all the metals. The order of the resistance of strain E1 to metals
Table 2 MIC (mmol/L) of various heavy metals
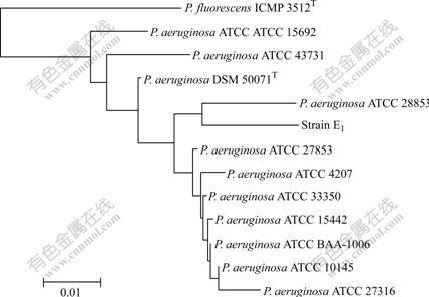
Fig.3 Phylogenetic tree derived from 16S rDNA sequence data of strain E1 (Scale bar corresponds to 1% nucleotide sequence difference)
is found to be Cd>Mn>Zn>Cu>Pb>Co. The resistance to heavy metals of strain E1 is far higher than that of E.coli.
3.3 Resistance to antibiotics
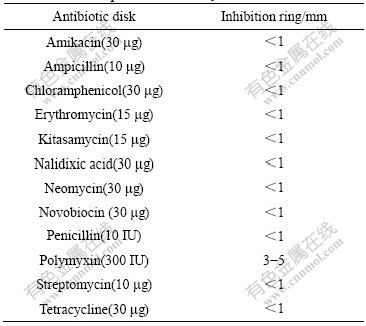
In order to determine the resistance to antibiotics, the reactions to 12 antibiotics were examined by the disk diffusion method. After incubating for 36-48 h, the plates with antibiotics disk appeared colonies. In the plates inoculated with strain E1, except that the plates with polymyxin have 3-5 mm inhibition ring, there is no obvious inhibition ring in the other plates (Table 3).
Table 3 Antibiotic profile of strain E1
Under environmental conditions of heavy metal stress, microorganisms might have developed various mechanisms to resist antibiotics and tolerate metals. Microbes surviving in polluted soils usually change intrinsic biochemical and structural properties, physiological, and genetic adaptation. Some metal- resistant microorganisms have been reported [23-25]. YILMAZ [23] isolated a heavy-metal-resistant bacterium Bacillus sp. EB1 from heavy-metal-contaminated soil. The strain can resist Cd, Ni, Cu, Co, Pb, Zn and antibiotics, and the Cd MIC is 2.5 mmol/L. HU et al [25] isolated two Cd-resistant bacteria identified as Bacillus cereus and Enterobacter cloacae. The two strains can also resist to Cu, Co, Pb, Zn and antibiotics, and the Cd MICs are separately 1 200 and 2 000 mg/L. Resistance to heavy metals and antibiotics of strain E1 were tested. In comparison to previous metal-resistant bacteria, strain E1 exhibits more tolerance to heavy metals and more resistance to the antibiotics, and the Cd MIC is 18.5 mmol/L (or 2 072 mg/L). Strain E1 is isolated from the melt soil with multiple heavy metals. The bacterium is exposed to the heavy metals for a long time, which might change its structure and function to adapt to heavy-metal-polluted environment, such as smaller size of bacteria, acquisition of resistance factors, and gene mutation. The high levels of resistance and the widespread tolerance to metals and antibiotics might be associated with its complex survival environment. The strain can grow in the heavy metals solution with the high level resistance to heavy metals. So it is possible to remove cadmium from wastewater containing cadmium.
3.4 Removal of cadmium
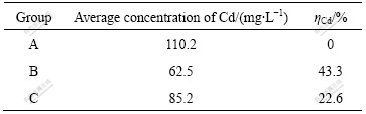
It can be found from Table 4 that, the average Cd concentrations of groups B and C are obviously lower than that of group A. From time and engineering cost, treating for 24 h is appropriate. Compared with group A without bacteria, the Cd concentration of group B with living bacteria decreases by 47.7 mg/L, and the average Cd concentration of group C with dead bacteria decreases by 25.0 mg/L. The result shows that the strain has good metal removal properties, both living and dead E1 can remove cadmium from solution.
Table 4 Cadmium removal of strain E1
The aim of studying the microorganism growth in heavy metal environment is to seek for the remediation measures. Many scholars have studied heavy metal removal from wastewater by microorganisms [13, 15, 20]. Generally, the process of microorganisms removal of heavy metals includes biosorption and bioaccumulation [26]. Biosorption process means metal ions are adsorbed by microorganisms through the biochemical reactions including complex, chelate, ions exchange, and adsorption. Both living and non-living cells have biosorption. Bioaccumulation happens only in living cell. It is an active process and needs energy provided by metabolism of microorganisms. KEFALA et al [27] reported that non-living bacteria were better than living bacteria in bioremediation because non-living cannot be affected by the metal toxicity. But TANGAROMSUK et al [28] reported that the metal removal efficiency of living cell was higher than that of non-living cell. In this work, the result is in agreement with the latter. The causes might be that the strain can resist Cd, and Cd has little influence on the stain. During the treatment process, living cell removes Cd by biosorption and bioaccumulation, while non-living cell removes Cd only by biosorption. So the Cd removal capacity of living cell is better than that of non-living cell.
4 Conclusions
(1) Strain E1 with great cadmium-resistance is isolated. It is Gram-negative, rod-shaped with polar flagellum, and the size is (0.5-1.0) μm×(2.0-3.0) μm. Incubated with Cd medium, the size becomes smaller. Based on the strain characterizations and the 16 rDNA sequence comparison, strain E1 is identified as P. aeruginosa.
(2) Besides the resistance to Cd, strain E1 is also tolerant to other metals such as Mn, Zn, Cu, Pb, Co on solid media plate. Strain E1 exhibits high resistance to wide antibiotic range.
(3) Both living and non-living cells of strain E1 can remove Cd from solution. After being treated for 24 h, Cd concentrations in the living cell and non-living cell, solution decrease by 47.7 mg/L (from 110.2 to 62.5 mg/L) and 25.0 mg/L (from 110.2 to 85.2 mg/L), and the corresponding cadmium removal rates reach 43.3% and 22.6%, respectively.
References
[1] DENG X, YI X E, LIU G. Cadmium removal from aqueous solution by gene-modified Escherichia coli JM109 [J]. Journal of Hazardous Materials, 2007, 139(2): 340-344.
[2] ABOU-SHANAB R A I, BERKUM P, ANGLE J S. Heavy metal resistance and genotypic analysis of metal resistance genes in gram-positive and gram-negative bacteria present in Ni-rich serpentine soil and in the rhizosphere of Alyssum murale [J]. Chemosphere, 2007, 68(2): 360-367.
[3] BASTA N T, GRADWOHL R, SNETHEN K L, SCHRODER J L. Chemical immobilization of lead, zinc, and cadmium in smelter-contaminated soils using biosolids and rock phosphate [J]. Journal Environment Quality, 2001, 30(4): 1222-1230.
[4] FILIPIC M, FATUR T, VUDRAG M. Molecular mechanisms of cadmium induced mutagenicity [J]. Human and Experimental Toxicology, 2006, 25(2): 67-77.
[5] GARY G, SCHWARTZ, ISILDINHA M. Is cadmium a cause of human pancreatic cancer? [J]. Cancer Epidemiol Biomarkers Prev, 2000, 9(2): 139-145.
[6] BABUK R, RAJMOHAN H R R, RAJANB K M, KARUNA K M. Plasma lipid peroxidation and erythrocyte antioxidant enzymes status in workers exposed to cadmium [J]. Toxicology and Industrial Health, 2006, 22(8): 329-335.
[7] YANG Z Q, YANG S F, QIAN S Y, HONG J S, KADIISKA M B, TENNANT R W, WAALKES M P, LIU J. Cadmium-induced toxicity in rat primary mid-brain neuroglia cultures: Role of oxidative stress from microglia [J]. Toxicol Sci, 2007, 98(2): 488-494.
[8] BRZ?SKA M M, MONIUSZKO-JAKONIUK J. Low-level exposure to cadmium during the lifetime increases the risk of osteoporosis and fractures of the lumbar spine in the elderly: Studies on a rat model of human environmental exposure [J]. Toxicol Sci, 2004, 82(2): 468-477.
[9] ANA N A, ELIZABETH S, SHARRETT A R, CALDERON E. Lead, cadmium, smoking, and increased risk of peripheral arterial disease [J]. Circulation, 2004, 109(6): 3196-3201.
[10] SHIN H J, LEE B H, YEO M G, OH S H, PARK J D, PARK K K. Induction of orphan nuclear receptor Nur77 gene expression and its role in cadmium-induced apoptosis in lung [J]. Carcinogenesis, 2004, 25(8): 1467-1475.
[11] GRASSESCHI R M, RAMASWAMY R B, LEVINE D J, KLAASSEN C D, WESSELIUS L J. Cadmium accumulation and detoxification by alveolar macrophages of cigarette smokers [J]. Chest, 2003, 124(5): 1924-1928.
[12] NISHIJO M, MORIKAWA Y, NAKAGAWA H, TAWARA K, MIURA K, KIDO T, IKAWA A, KOBAYASHI E, NOGAWA K. Causes of death and renal tubular dysfunction in residents exposed to cadmium in the environment [J]. Occup Environ Med, 2006, 63(8): 545-550.
[13] ZIAGOVA M, DIMITRIADIS G, ASLANIDOU D, PAPAIOANNOU X, LITOPOULOU T E, LIAKOPOULOU K M. Comparative study of Cd(II) and Cr(VI) biosorption on Staphylococcus xylosus and Pseudomonas sp. in single and binary mixtures[J]. Biosensors Technol, 2007, 98(15): 2859-2865.
[14] QIN Wen-qing, LAN Zhuo-yue, LI Wei-zhong. Recovery of zinc from low-grade zinc oxide ores by solvent extraction [J]. Journal of Central South University of Technology, 2003, 10 (2): 98-102.
[15] YU Jun-xia, TONG Mi, SUN Xiao-mei, LI Bu-hai. Biomass grafted with polyamic acid for enhancement of cadmium (II) and lead (II) biosorption [J]. Biochemical Engineering Journal, 2007, 33(2): 126-133.
[16] HINOJOSA M B, CARREIRA J A, ROBERTO G R, RICHARD P D. Microbial response to heavy metal-polluted soils: Community analysis from phospholipid-linked fatty acids and ester-linked fatty acids extracts [J]. J Environ Qual, 2005, 34(5): 1789-1800.
[17] ZOUBOULIS A I, LOUKIDOU M X, MATIS K A. Biosorption of toxic metals from aqueous solutions by bacteria strains isolated from metal-polluted soils [J]. Process Biochemistry, 2004, 39(8): 909-916.
[18] OZDEMIR G, CEYHAN N, OZTURK T, AKIRMAK F, COSAR T. Biosorption of chromium (VI), cadmium (II) and copper (II) by Pantoea sp. TEM18 [J]. Chemical Engineering Journal, 2004, 102(3): 249-253.
[19] LEBEAU T, BAGOT D, J?Z?QUEL B, FABRE B. Cadmium biosorption by free and immobilised microorganisms cultivated in a liquid soil extract medium: Effects of Cd, pH and techniques of culture [J]. The Science of the Total Environment, 2002, 291(1/3): 73-83.
[20] YUXia, CHAILi-yuan, MINXiao-bo. Removal of lead in wastewater by immobilized inactivated cells of rhizopus oligosporus [J]. Journal of Central South University of Technology, 2003, 10(4): 313-317.
[21] DONG Xiu-zhu, CAI Miao-ying. Manual of common determinative bacteriology [M]. Beijing: Science Press, 2001. (in Chinese)
[22] ZHOU Hong-bo, ZENG Xiao-xi, IU Fei-fei, HU Yue-hua. Screening, identification and desilication of a silicate bacteria [J]. Journal of Central South University of Technology, 2006, 13(4): 337-341.
[23] YILMAZ E I. Metal tolerance and biosorption capacity of Bacillus circulars strain EB1 [J]. Research in Microbiology, 2003, 154(6): 409-415.
[24] VERMA T, SRINATH T, GADPAYLE R U, RAMTEKE P W, HANS R K, GARG S K. Chromate tolerant bacteria isolated from tannery effluent [J]. Bioresource Technol, 2001, 78(1): 31-35.
[25] HU Qing, DOU Min-na, QI Hong-yan, XIE Xiang-ming, ZHUANG Guo-qiang, YANG Min. Detection isolation, and identification of cadmium-resistant bacteria based on PCR-DGGE [J]. Journal of Environmental Sciences, 2007, 19(9): 1114-1119.
[26] GUPTA G, KEEGAN B. Bioaccumulation and biosorption of lead by poultry litter microorganisms [J]. Poultry Science, 1998, 77(3): 400-404.
[27] KEFALA M I, ZOUBOULIS A I, MATIS K A. Biosorption of cadmium ions by actinomycetes and separation by flotation [J]. Environmental Pollution, 1999, 104(2): 283-293.
[28] TANGAROMSUK J, POKETHITIYOOK P, KRUATRACHUE M, UPATHAM E S. Cadmium biosorption by sphingomonas paucimobilis biomass [J]. Bioresource Technology, 2002, 85(1): 103-105.
(Edited by YANG You-ping)
Foundation item: Project (50621063) supported by the National Natural Science Foundation of China; Project (2004CB619204) supported by the Major State Basic Research and Development Program of China
Received date: 2008-07-23; Accepted date: 2008-09-26
Corresponding author: LIU Xue-duan, Professor; Tel: +86-731-8836019; E-mail: xueduanliu@sina.com